|
Extraction and Conversion of Protein Sequence
Download the Hemoglobin PDB file used in this example by clicking here. Save the file as 1A00_pdb.txt.
Open Vect and open the 1A00_pdb.txt file through the 'Open' icon or select Files > Open files from the pull down menu. The file should appear in the body. Use the scroll bar to view the whole file. The number at the left most side represent line numbers (1, 2, 3, …) and the other numbers (1, 1, 1, …) besides the line numbers represent the level of data.
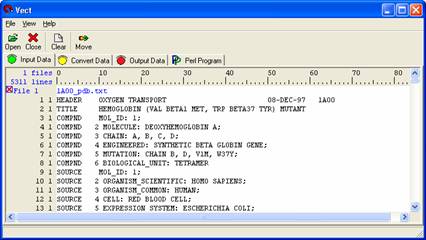
We will first be selecting the protein sequence located at the bottom of the Hemoglobin PDB File.
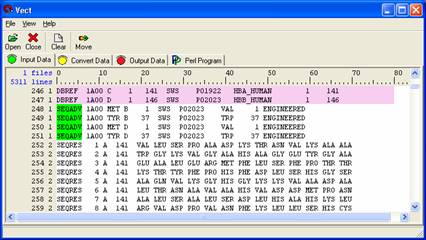
Right click on 'SEQADV' located in field 0, line 251 and select New Block Open Condition from the pull down menu. A green highlighted box will appear and the level from the origin will change because you open a new block. . Right click on ‘SEQADV’ and choose ‘Selection Exclusive’ so that data in this line would not be included when you select data you want.
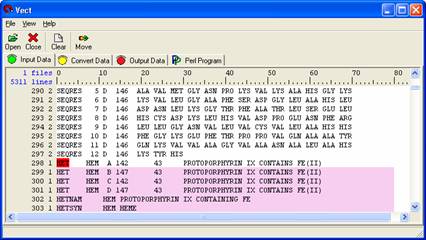
Scroll down to the bottom and right click on 'HET' located in field 0, line 298, choose New Block Close Condition from the pull down menu. A red highlighted box will appear and levels after ‘HET” will change to 1 because you close new block. Right click on ‘HET’ and choose ‘Selection Exclusive’ so that data in this line would not be included when you select data you want.
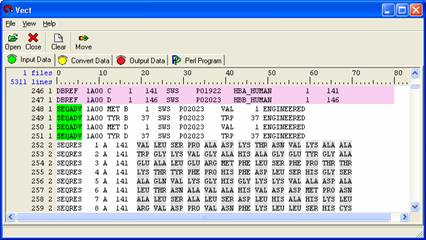
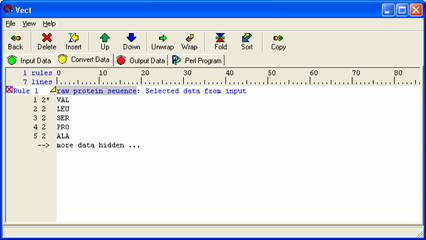
Left click on the first column containing the protein sequence. This will highlight the entire column grey, meaning it is selected. Then left click on the second column, then the third column, and so forth until you have selected all thirteen of them. You are making a column selection and fields one through thirteen should be highlighted in grey. The figure below shows the final appearance of the protein sequence selections.
Note: If you do not have any grey highlighted regions in the 'Input Data' panel then you have not selected any data and no data can be moved over to the next panel.
Select 'Move' from the icon panel and give your rule a descriptive name. Here you just to type “raw protein sequence” and press enter key to follow the remaining tutorial. This moves the data that you have selected to the 'Convert Data' panel where rule-dependent rules can be incorporated for better data extraction. (Rules are just a bunch of conditions you set so that you can extract data).
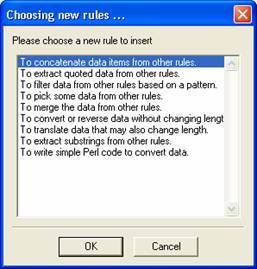
Select 'Insert' from the icon panel and select the Concatenation Rule. Give your rule a descriptive name. Here just type “protein sequence” for example

|